OptZyme: In Silico Enzyme Redesign Using Transition State Analogues
ID# 2013-4127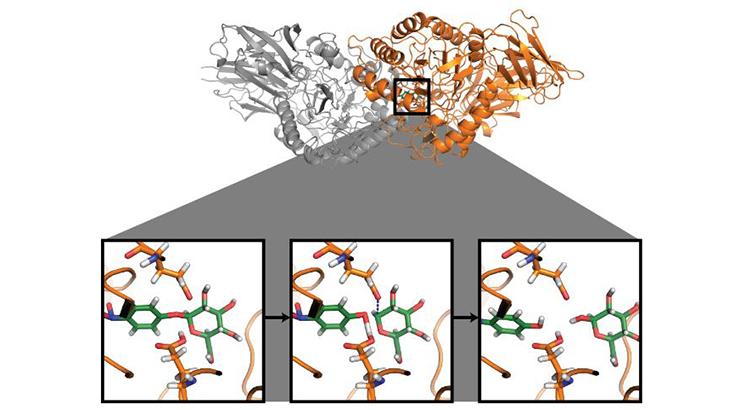
Technology Summary
OptZyme is software used to discover promising enzyme redesigns that can be experimentally tested. OptZyme does not use complete quantum mechanics analysis making it less computationally demanding. OptZyme is instead predicated on derived equations. OptZyme utilizes known transition state analogue (TSA) structures as proxies for unknown rate-limiting transition state (TS) structures. TSAs interfere with enzyme catalytic activity by mimicking the geometry of TSs and preferentially binding with the enzyme over the substrate, preventing the reaction from proceeding. The inventors have shown that mutations that minimize the interaction energy (IE) of the enzyme with its TSA, rather than with its substrate, allows for the identification of lower transition state formation energy barrier.
Application & Market Utility
OptZyme helps design new commercial enzymes, providing strong evidence before using experimental approaches. OptZyme is suited for systems where solute entropy change upon binding is assumed to be negligible relative to other terms, substrate binding is not a consequence of “induced fit”, and equilibrium following the rate-limiting step strongly favors product release. OptZyme constructs a library of mutants with improved enzyme catalytic parameters for a similar substrate by identifying novel contacts with the ligand that were absent from other established libraries.
Next Steps
Seeking prospective licensees; demo is available at http://maranas.che.psu.edu/submission/OptZyme.htm

